Similarity
Arguments
- object
A \(m \times p\)
numericmatrixordata.frameof count data (absolute frequencies giving the number of individuals for each category, i.e. a contingency table). Adata.framewill be coerced to anumericmatrixviadata.matrix().- ...
Currently not used.
- method
A
characterstring specifying the method to be used (see details). Any unambiguous substring can be given.
Value
A stats::dist object.
Details
\(\beta\)-diversity can be measured by addressing similarity between pairs of samples/cases.
bray, jaccard, morisita and sorensen indices provide a scale of
similarity from \(0\)-\(1\) where \(1\) is perfect similarity and
\(0\) is no similarity.
brainerd is scaled between \(0\) and \(200\).
brainerdbrayBray-Curtis similarity (a.k.a. Dice-Sorensen quantitative index).
jaccardmorisitasorensen
For jaccard and sorensen, data are standardized on a presence/absence
scale (\(0\)/\(1\)) beforehand.
References
Magurran, A. E. (1988). Ecological Diversity and its Measurement. Princeton, NJ: Princeton University Press. doi:10.1007/978-94-015-7358-0 .
See also
index_binomial(), index_brainerd(), index_bray(),
index_jaccard(), index_morisita(), index_sorensen()
Other diversity measures:
diversity(),
evenness(),
heterogeneity(),
occurrence(),
plot.DiversityIndex(),
plot.RarefactionIndex(),
profiles(),
rarefaction(),
richness(),
she(),
simulate(),
turnover()
Examples
## Data from Huntley 2004, 2008
data("pueblo")
## Brainerd-Robinson measure
(C <- similarity(pueblo, "brainerd"))
#> Atsinna Cienega Mirabal PdMuertos Hesh LowPesc BoxS
#> Cienega 164.36782
#> Mirabal 152.38095 138.09524
#> PdMuertos 179.31034 150.57471 152.38095
#> Hesh 82.75862 55.55556 66.66667 103.44828
#> LowPesc 89.21023 62.00717 73.11828 109.89989 193.54839
#> BoxS 82.75862 55.55556 114.28571 102.17114 103.70370 103.70370
#> Ojo Bon 27.58621 22.22222 19.04762 20.68966 26.66667 25.80645 22.22222
#> S170 27.58621 22.22222 19.04762 20.68966 26.66667 25.80645 22.22222
#> Ojo Bon
#> Cienega
#> Mirabal
#> PdMuertos
#> Hesh
#> LowPesc
#> BoxS
#> Ojo Bon
#> S170 190.53030
plot_spot(C)
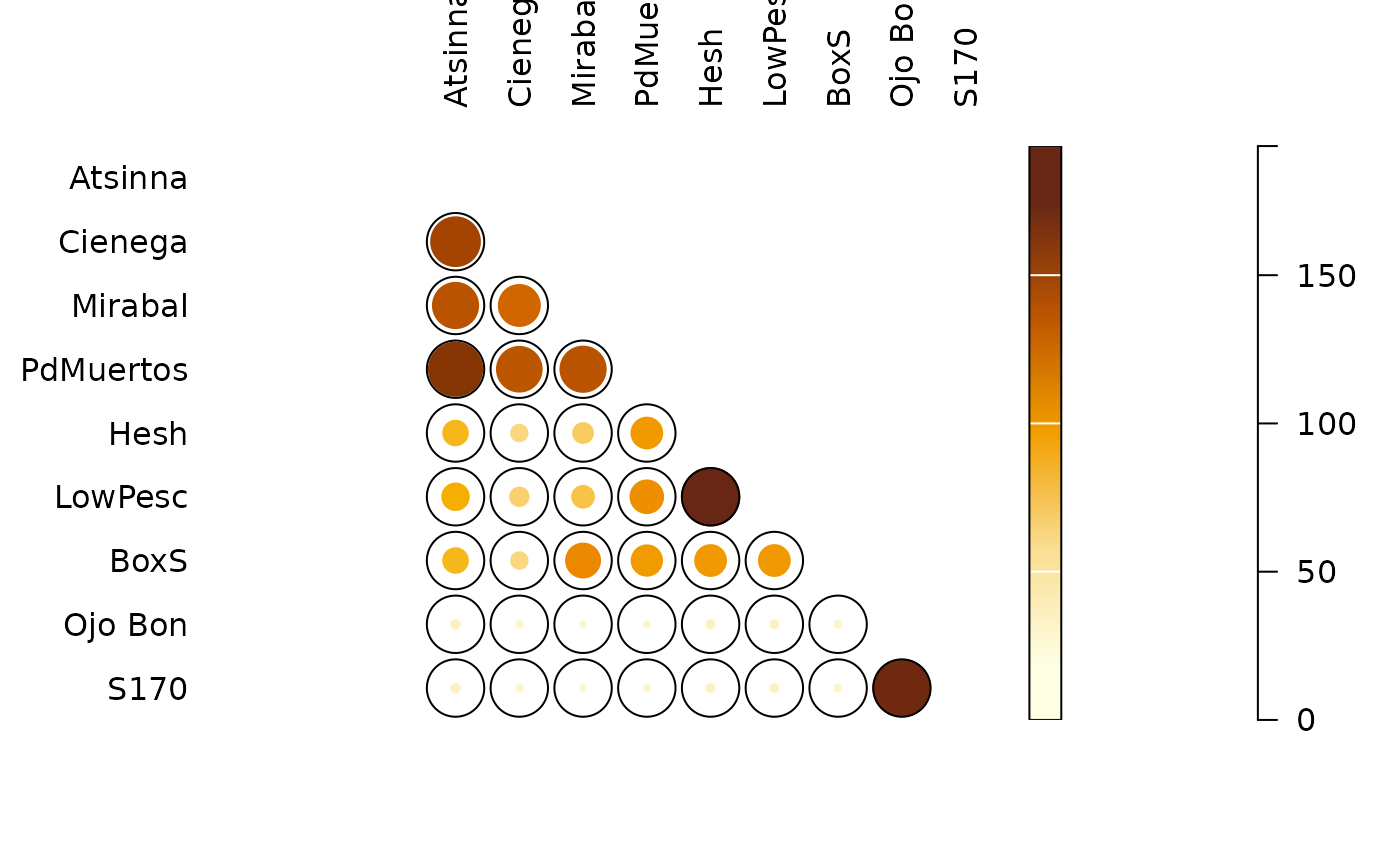 ## Data from Magurran 1988, p. 166
data("aves")
## Jaccard measure (presence/absence data)
similarity(aves, "jaccard") # 0.46
#> unmanaged
#> managed 0.4615385
# Bray and Curtis modified version of the Sorensen index (count data)
(sim <- similarity(aves, "bray")) # 0.44
#> unmanaged
#> managed 0.4442754
# Bray and Curtis dissimilarity
1 - sim
#> unmanaged
#> managed 0.5557246
## Data from Magurran 1988, p. 166
data("aves")
## Jaccard measure (presence/absence data)
similarity(aves, "jaccard") # 0.46
#> unmanaged
#> managed 0.4615385
# Bray and Curtis modified version of the Sorensen index (count data)
(sim <- similarity(aves, "bray")) # 0.44
#> unmanaged
#> managed 0.4442754
# Bray and Curtis dissimilarity
1 - sim
#> unmanaged
#> managed 0.5557246
