Computes the distances between all rows of in x.
Usage
# S4 method for class 'CompositionMatrix'
dist(x, method = "euclidean", diag = FALSE, upper = FALSE, p = 2)Arguments
- x
A
CompositionMatrixobject.- method
A
characterstring specifying the distance measure to be used. Seestats::dist()for the available distances.- diag
A
logicalscalar indicating whether the diagonal of the distance matrix should be printed.- upper
A
logicalscalar indicating whether the upper triangle of the distance matrix should be printed.- p
An
integergiving the power of the Minkowski distance.
Value
A stats::dist object.
Details
Distances are computed on CLR-transformed data.
References
Aitchison, J. (1986). The Statistical Analysis of Compositional Data. London: Chapman and Hall, p. 64-91.
Greenacre, M. J. (2019). Compositional Data Analysis in Practice. Boca Raton: CRC Press.
See also
Other statistics:
aggregate(),
condense(),
covariance(),
mahalanobis(),
margin(),
mean(),
pip(),
quantile(),
scale(),
variance(),
variance_total(),
variation()
Examples
## Data from Aitchison 1986
data("hongite")
## Coerce to compositional data
coda <- as_composition(hongite)
## Aitchison distance
## (euclidean distance between CLR-transformed compositions)
d <- dist(coda)
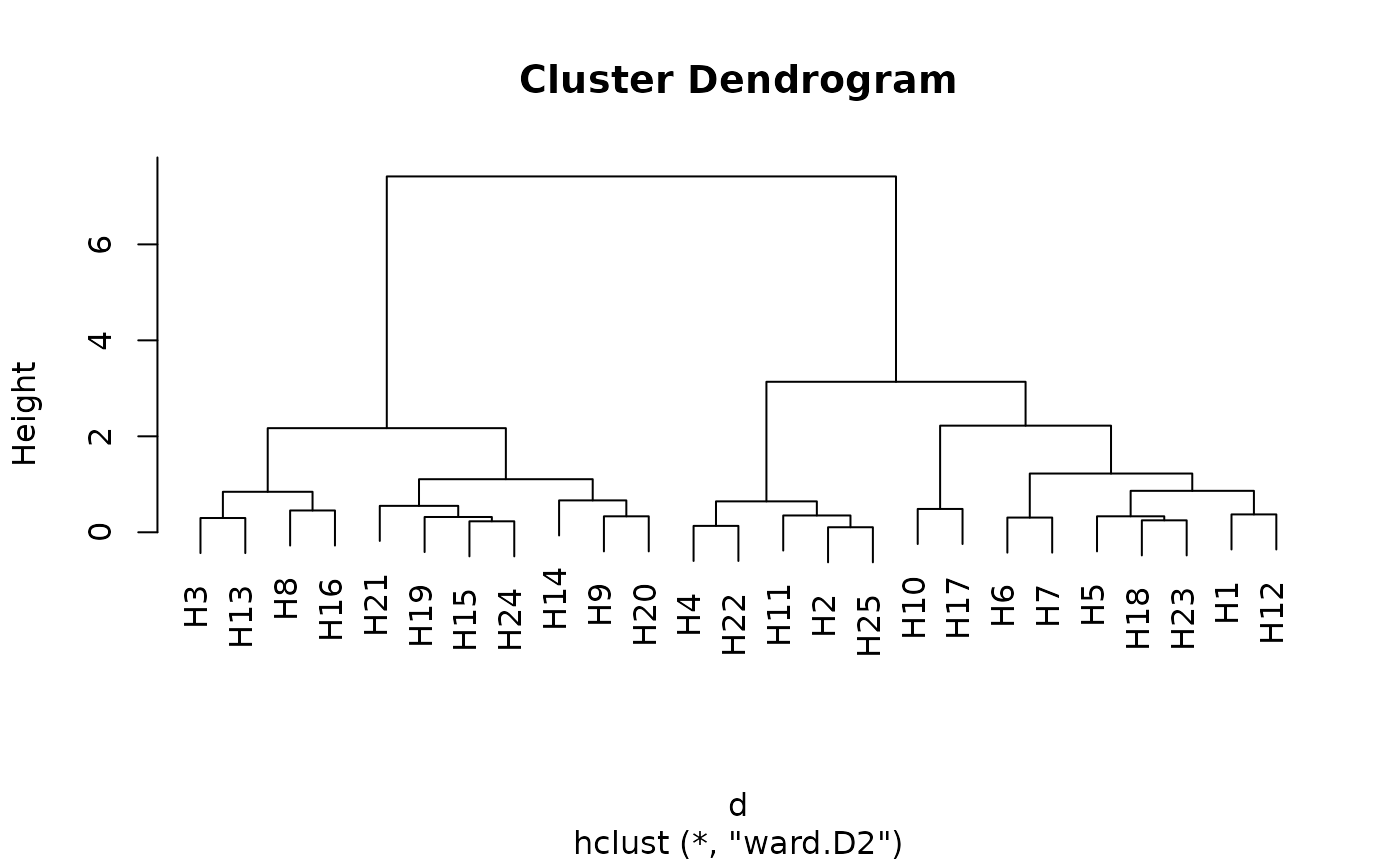
## Cluster dendrogram
h <- hclust(d, method = "ward.D2")
plot(h)