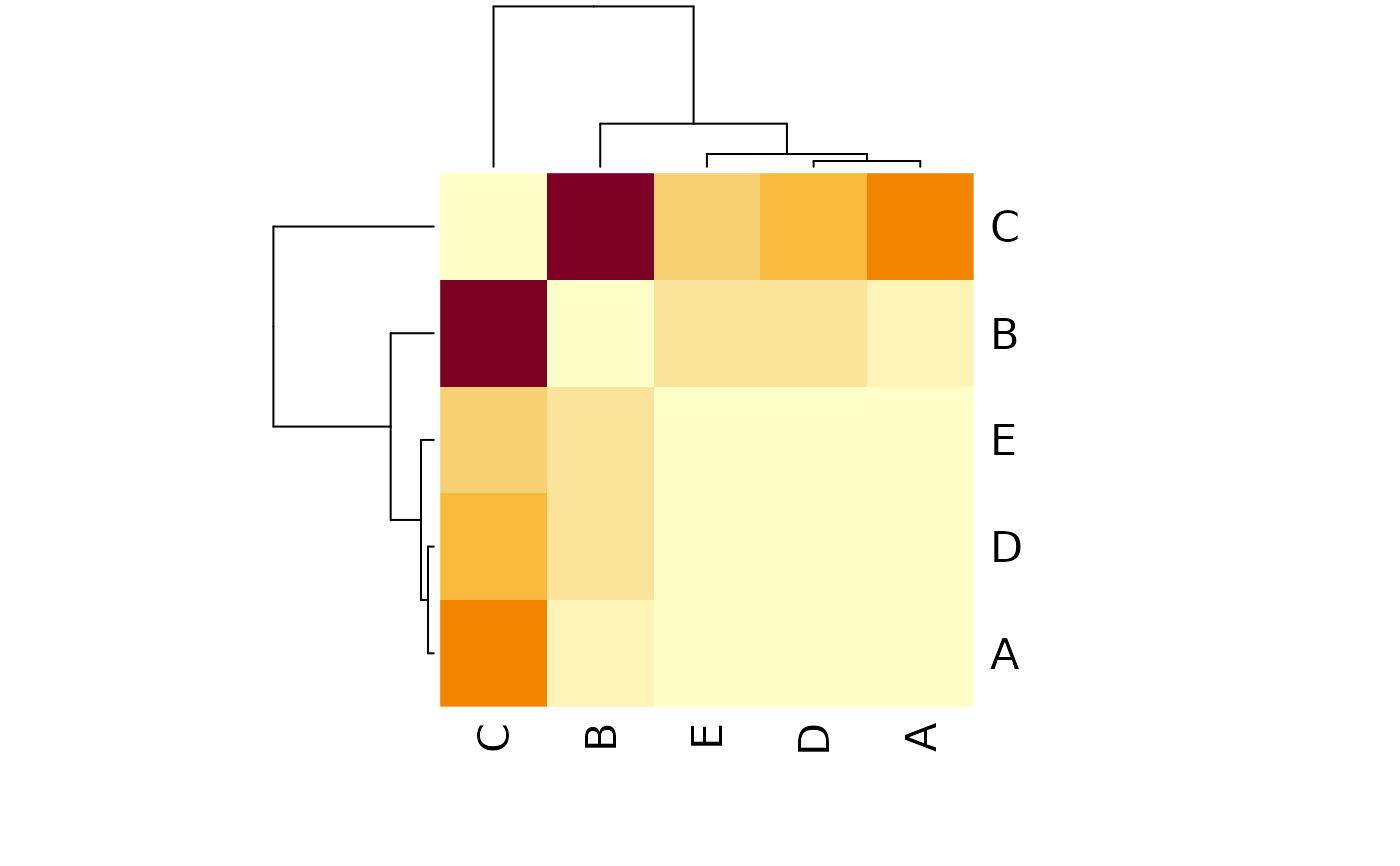Computes an index of association between parts.
Arguments
- x
A
CompositionMatrixobject.- ...
Currently not used.
Value
A matrix.
Details
The proportionality index of parts (PIP) is based on the variation matrix, but maintains the range of values whithin \((0,1)\).
References
Egozcue, J. J.. & Pawlowsky-Glahn, V. (2023). Subcompositional Coherence and and a Novel Proportionality Index of Parts. SORT, 47(2): 229-244. doi:10.57645/20.8080.02.7 .
See also
Other statistics:
aggregate(),
condense(),
covariance(),
dist,
mahalanobis(),
margin(),
mean(),
quantile(),
scale(),
variance(),
variance_total(),
variation()
Examples
## Data from Aitchison 1986
data("hongite")
## Coerce to compositional data
coda <- as_composition(hongite)
## Variation matrix
## (Aitchison 1986, definition 4.4)
(varia <- variation(coda))
#> A B C D E
#> A 0.00000000 0.2592742 1.5328586 0.08281464 0.1385604
#> B 0.25927416 0.0000000 3.0006804 0.54727146 0.6490135
#> C 1.53285862 3.0006804 0.0000000 1.11145146 0.9476375
#> D 0.08281464 0.5472715 1.1114515 0.00000000 0.1870579
#> E 0.13856035 0.6490135 0.9476375 0.18705791 0.0000000
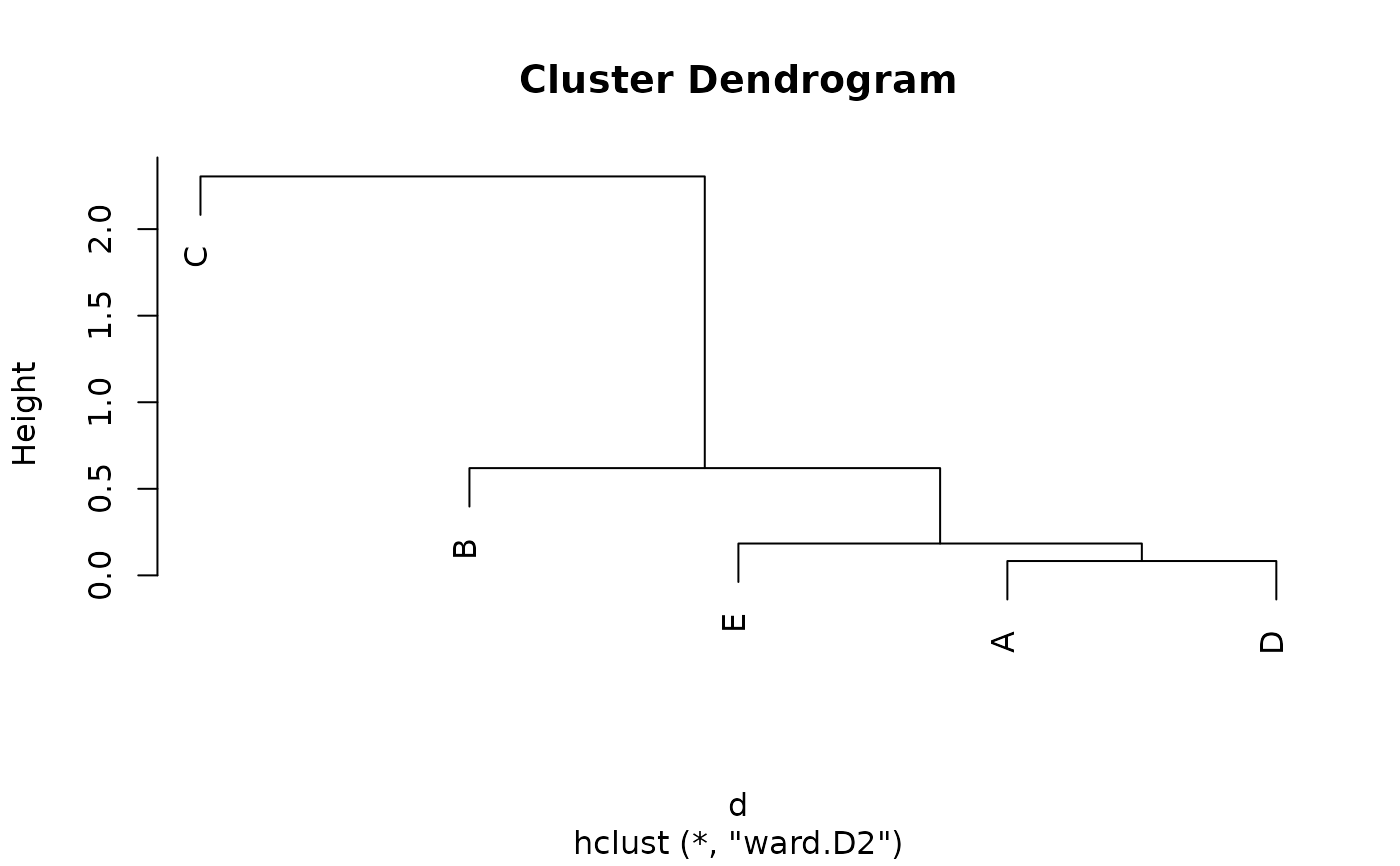
## Cluster dendrogram
d <- as.dist(varia)
h <- hclust(d, method = "ward.D2")
plot(h)
 ## Heatmap
stats::heatmap(
varia,
distfun = stats::as.dist,
hclustfun = function(x) stats::hclust(x, method = "ward.D2"),
symm = TRUE,
scale = "none"
)
## Heatmap
stats::heatmap(
varia,
distfun = stats::as.dist,
hclustfun = function(x) stats::hclust(x, method = "ward.D2"),
symm = TRUE,
scale = "none"
)