Computes a principal components analysis based on the singular value decomposition.
Usage
# S4 method for class 'CompositionMatrix'
pca(
object,
center = TRUE,
scale = FALSE,
rank = NULL,
sup_row = NULL,
sup_col = NULL,
weight_row = NULL,
weight_col = NULL
)
# S4 method for class 'LogRatio'
pca(
object,
center = TRUE,
scale = FALSE,
rank = NULL,
sup_row = NULL,
sup_col = NULL,
weight_row = NULL,
weight_col = NULL
)Arguments
- object
A
CompositionMatrixorLogRatioobject.- center
A
logicalscalar: should the variables be shifted to be zero centered?- scale
A
logicalscalar: should the variables be scaled to unit variance?- rank
An
integervalue specifying the maximal number of components to be kept in the results. IfNULL(the default), \(p - 1\) components will be returned.- sup_row
A
vectorspecifying the indices of the supplementary rows.- sup_col
A
vectorspecifying the indices of the supplementary columns.- weight_row
A
numericvector specifying the active row (individual) weights. IfNULL(the default), uniform weights are used. Row weights are internally normalized to sum 1- weight_col
A
numericvector specifying the active column (variable) weights. IfNULL(the default), uniform weights (1) are used.
Value
A dimensio::PCA object. See dimensio::pca() for details.
Methods (by class)
pca(CompositionMatrix): PCA of centered log-ratio, i.e. log-ratio analysis (LRA).
References
Aitchison, J. and Greenacre, M. (2002). Biplots of compositional data. Journal of the Royal Statistical Society: Series C (Applied Statistics), 51: 375-392. doi:10.1111/1467-9876.00275 .
Filzmoser, P., Hron, K. and Reimann, C. (2009). Principal component analysis for compositional data with outliers. Environmetrics, 20: 621-632. doi:10.1002/env.966 .
Examples
## Data from Day et al. 2011
data("kommos", package = "folio") # Coerce to compositional data
kommos <- remove_NA(kommos, margin = 1) # Remove cases with missing values
coda <- as_composition(kommos, groups = 1) # Use ceramic types for grouping
## Log-Ratio Analysis
X <- pca(coda)
#> PCA of centered log-ratio (CLR).
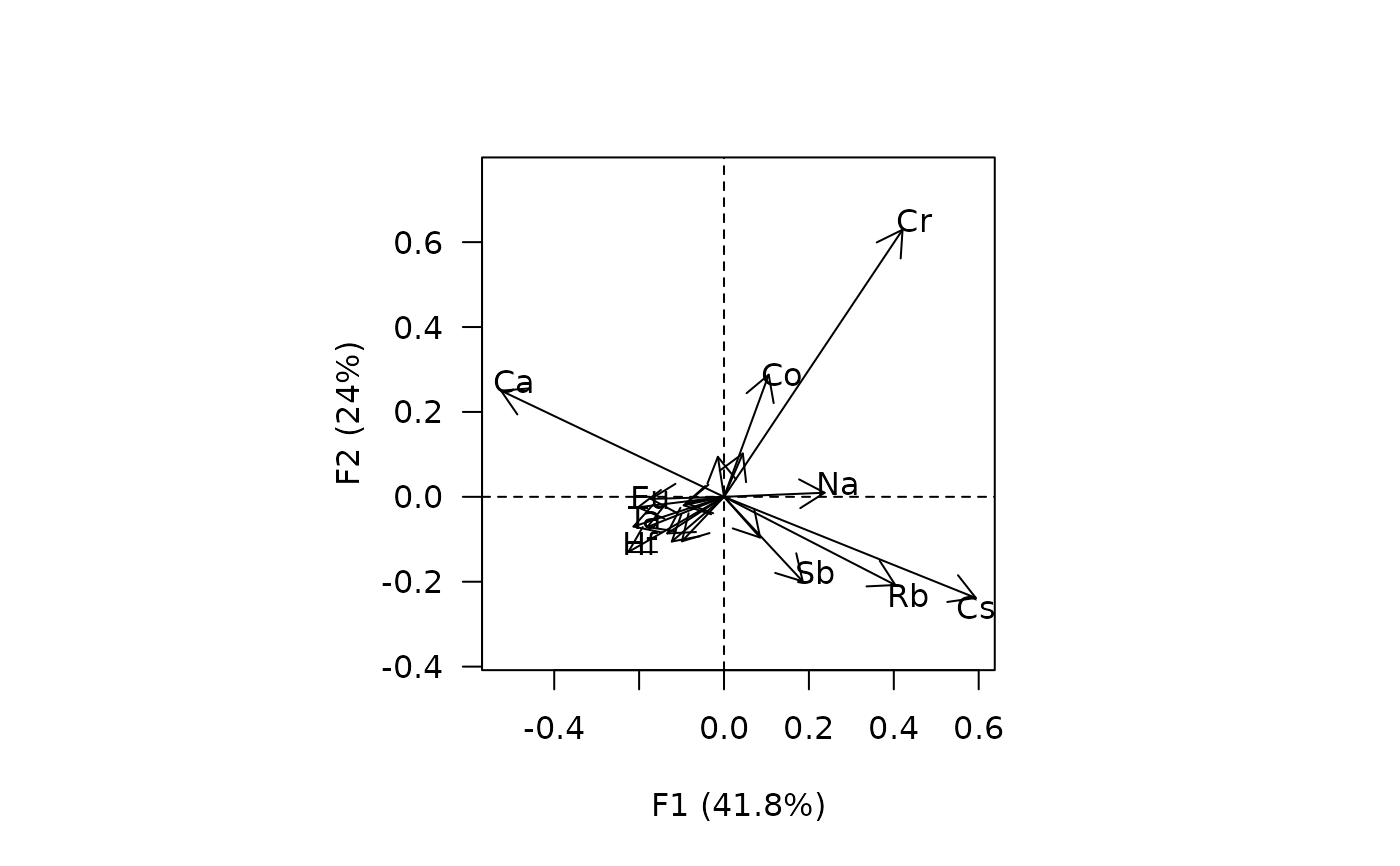
## Biplot
biplot(X)
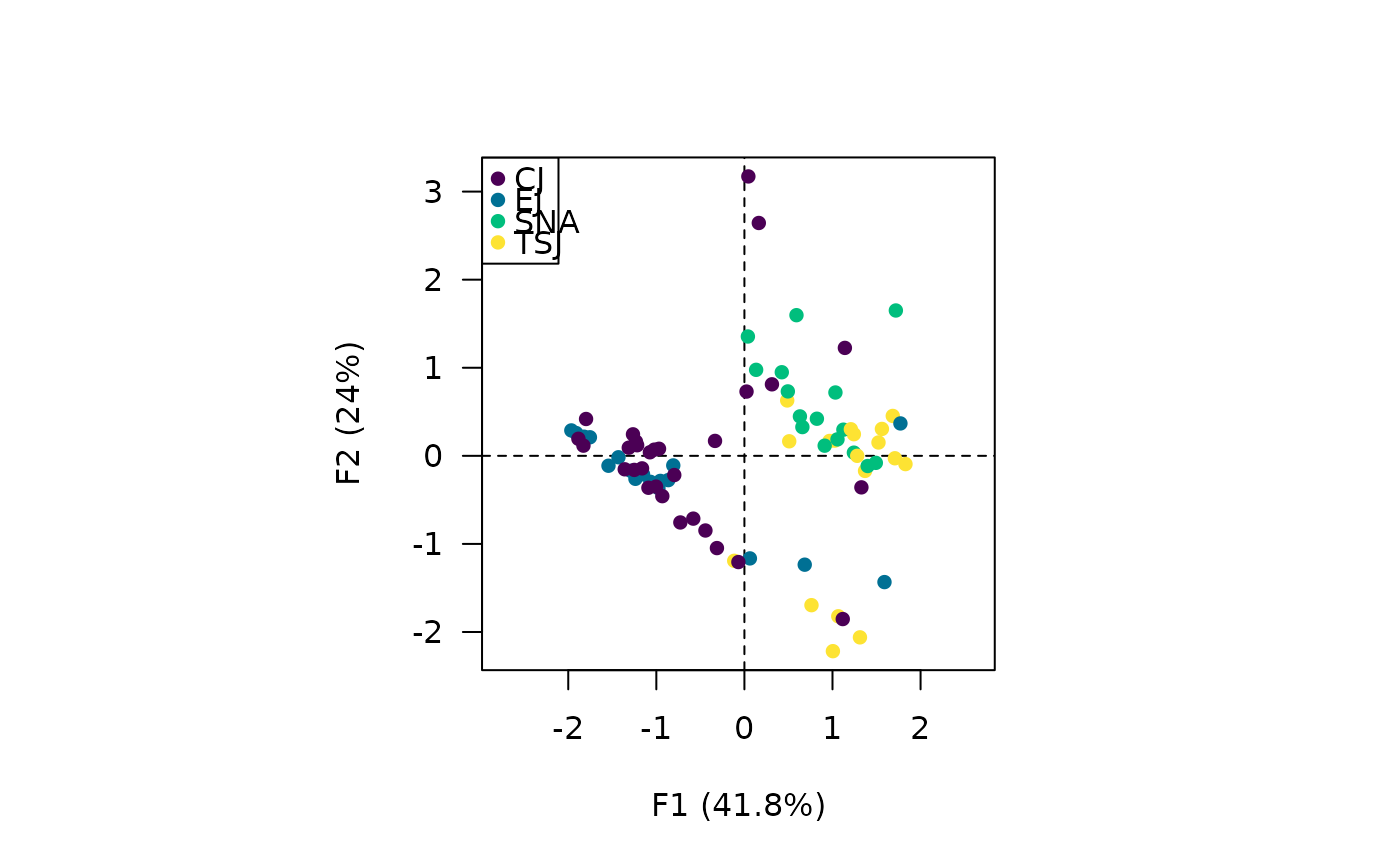 ## Explore results
viz_individuals(X, extra_quali = group_names(coda))
## Explore results
viz_individuals(X, extra_quali = group_names(coda))
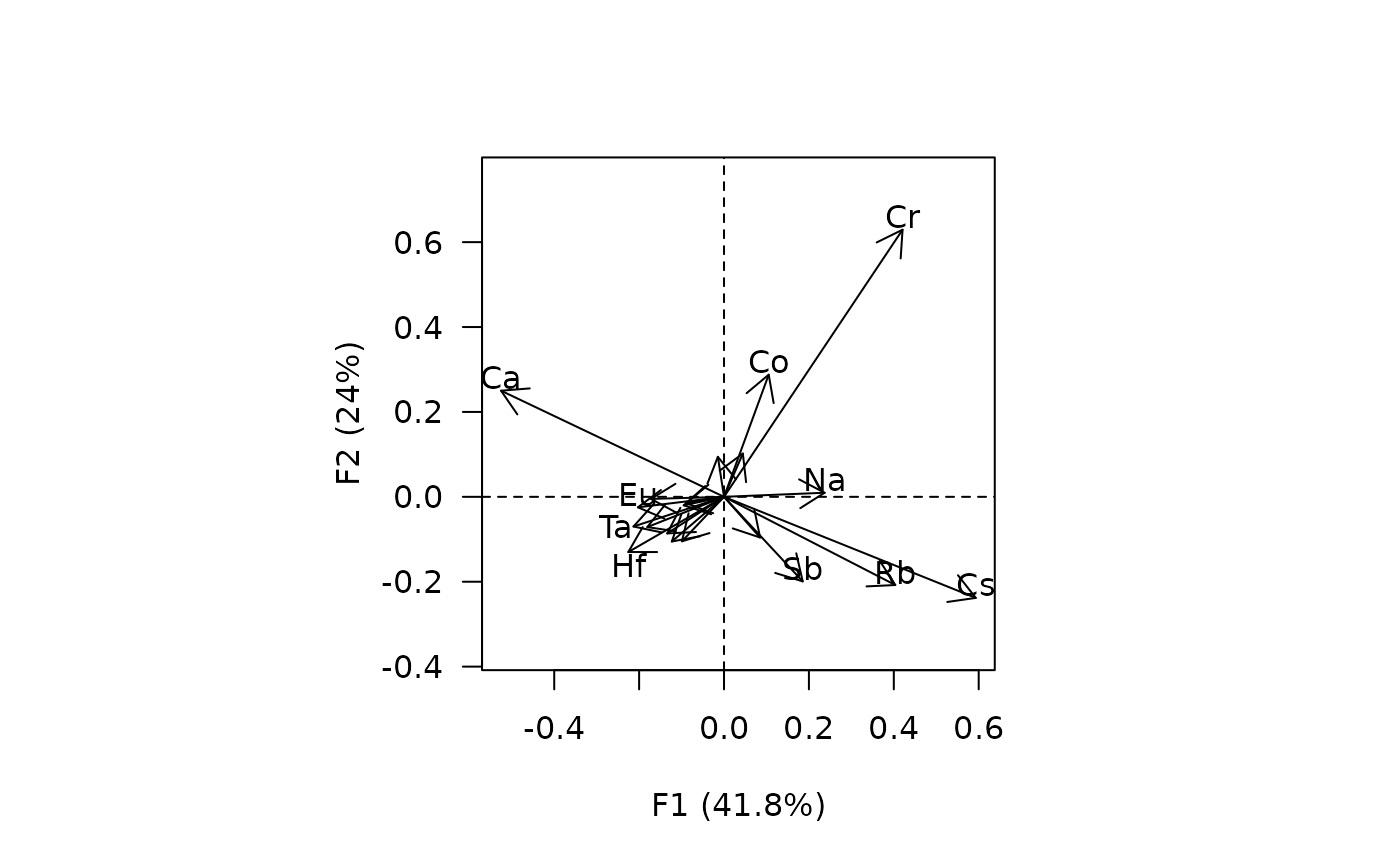 viz_variables(X)
viz_variables(X)